Linkage versus association--连锁与关联的区别---reprinted
文章目录
很多人在做关联分析,很多人在做连锁分析,其原理都是从遗传上找与表型相关marker。但两者是有区别的,特意转载下图,看到这幅图,你一定会对连锁和关联的区别有清楚的认识。
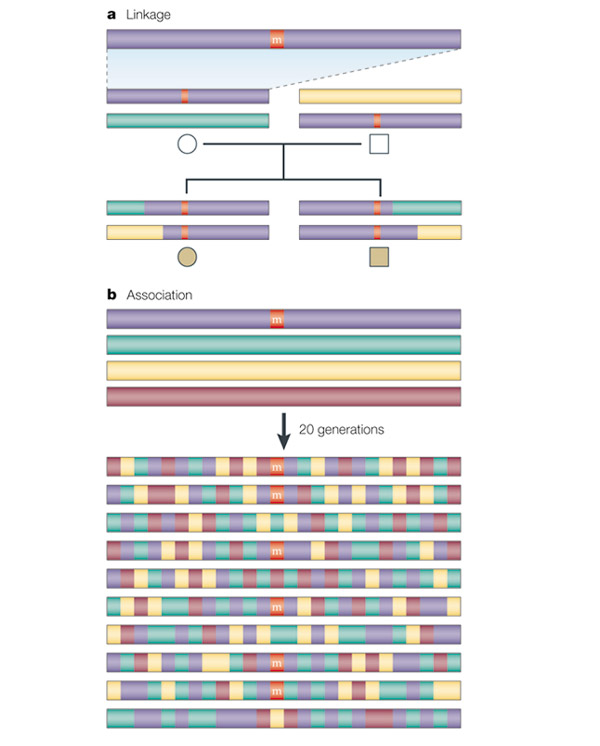
At a fundamental level, genetic association and linkage analysis rely on similar principles and assumptions. Both rely on the co-inheritance of adjacent DNA variants, with linkage capitalizing on this by identifying haplotypes that are inherited intact over several generations (such as in families or pedigrees of known ancestry), and association relying on the retention of adjacent DNA variants over many generations (in historic ancestries). Thus, association studies can be regarded as very large linkage studies of unobserved, hypothetical pedigrees. In growing populations, such as humans, recombination is the primary force that eliminates linkage and association over generations. When a functional mutation occurs (’m' in the figure) - perhaps one that contributes to disease - it does so on a haplotype of other pre-existing DNA variants. Because linkage focuses only on recent, usually observable ancestry, in whom there have been relatively few opportunities for recombination to occur, disease gene regions that are identified by linkage will often be large, and can encompass hundreds or even thousands of possible genes across many megabases of DNA (figure panel a). By contrast, association studies draw from historic recombination so disease-associated regions are (theoretically) extremely small in outbred random mating populations, encompassing only one gene or gene fragment (figure panel b). Through subsequent generations, as the disease mutation is transmitted, recombination will cause it to be separated from the specific alleles of its original haplotype. Particular DNA variants can remain together on ancestral haplotypes for many generations. This type of non-random association of alleles is known as linkage disequilibrium. It is linkage disequilibrium that provides the genetic basis for most association strategies.
Please note, the figure and contents are reprinted from Nature (title:Association study designs for complex diseases, author Lon R. Cardon & John I. Bell)
文章作者 zzx
上次更新 2016-01-11
